Dear Martin,
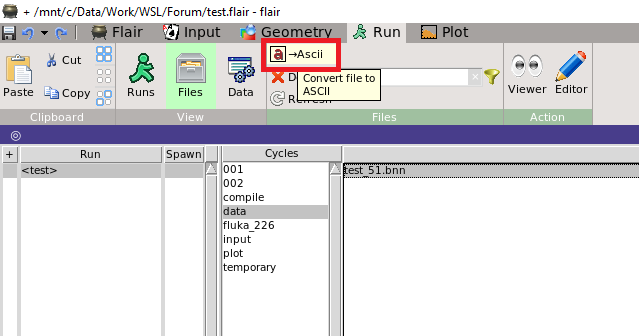
in addition to @cesc’s answer. You can convert a processed binary result to a readable text file with Flair as well, using the -> ASCII buttion on the Run/Files tab, as shown on the picture below.
Cheers,
David
Dear Martin,
in addition to @cesc’s answer. You can convert a processed binary result to a readable text file with Flair as well, using the -> ASCII buttion on the Run/Files tab, as shown on the picture below.

Cheers,
David