Dear FLUKA Experts,
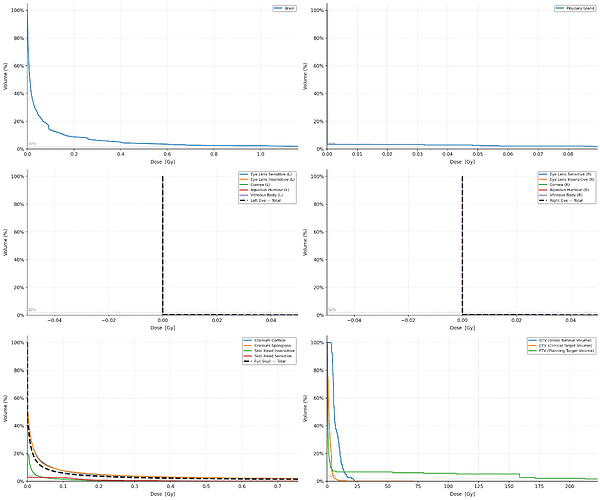
I am working on ICRP-145 mesh phantom, I need to calculate the Dose-Volume Histogram (DVH) for several organs, including the Brain, Eye Lens, and Pituitary Gland.
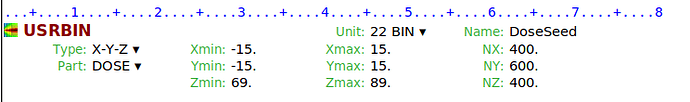
Following previous discussions on the forum ( How to score DVH in mathematical phantom ), I understand the standard workflow is to score dose in a 3D grid using a Cart USRBIN. I have two specific questions :
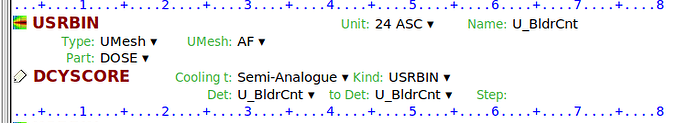
1- Since ICRP-145 is a mesh phantom, how can I precisely identify which voxel in my Cart USRBIN belongs to which specific organ region?
2- When defining the X/Y/Z boundaries of the USRBIN, should the grid cover the entire volume containing all organs of interest, or should a separate USRBIN be defined for each organ? :
From my .log file, I extracted the following geometric information for some organs (region ID, density, volume, and bounding box):
> Organ FLUKA_Region AM_ID Density(g/cm³) Volume(cm³) Mass(g) Xmin Xmax Ymin Ymax Zmin Zmax
> Brain 83 AM6100 1.041 1457.63 1517.3 -7.117 6.760 -8.742 8.890 73.183 86.548
> Pituitary_gland 158 AM11400 1.031 0.60312 0.622 -0.226 0.867 -2.469 -1.142 76.391 78.026
> Left_eye_components
> Eye_lens_sensitive(L) 88 AM6600 1.060 0.03669 0.0389 3.048 4.048 -9.049 -8.897 77.186 78.186
> Eye_lens_insensitive(L) 89 AM6601 1.060 0.17856 0.1893 3.048 4.048 -9.049 -8.629 77.186 78.186
> Cornea(L) 90 AM6700 1.087 1.0121 1.100 2.338 4.758 -9.384 -6.845 76.476 78.896
> Aqueous_humour(L) 91 AM6701 1.014 0.30007 0.3043 2.759 4.337 -9.324 -8.897 76.896 78.475
> Vitreous_body(L) 92 AM6702 1.019 5.9379 6.052 2.395 4.701 -8.893 -6.899 76.533 78.839
> Right_eye_components
> Eye_lens_sensitive(R) 93 AM6800 1.060 0.03669 0.0389 -3.277 -2.277 -9.243 -9.091 77.165 78.165
> Eye_lens_insensitive(R) 94 AM6801 1.060 0.17856 0.1893 -3.277 -2.277 -9.243 -8.823 77.165 78.165
> Cornea\_(R) 95 AM6900 1.087 1.0121 1.100 -3.987 -1.567 -9.578 -7.039 76.455 78.875
> Aqueous_humour(R) 96 AM6901 1.014 0.30007 0.3043 -3.566 -1.988 -9.518 -9.091 76.876 78.454
> Vitreous_body(R) 97 AM6902 1.019 5.9379 6.052 -3.930 -1.624 -9.088 -7.093 76.512 78.818
> Skull
> Cranium_cortical 50 AM2600 1.904 298.57 568.6 -7.537 7.182 -10.916 9.623 72.426 87.432
> Cranium_spongiosa 51 AM2700 1.252 327.96 410.6 -7.277 6.891 -9.930 9.310 72.585 87.259
> Skull_skin
> Skin_head_insensitive 166 AM12200 1.088 238.04 259.0 -8.895 9.578 -13.603 10.623 62.329 88.145
> Skin_head_sensitive 167 AM12201 1.088 7.7778 8.46 -8.890 9.577 -13.598 10.618 62.336 88.139